The host microbiome comprises the entirety of microorganisms colonizing a host. It is divided into different subpopulations that are related to certain body regions and organs. These subgroups are called microbiomes of these regions and organs.
There are differences in composition and tasks of the microbiomes of distinct hosts and organs. For example, the functional rumen microbiome in cattle is composed of a consortium of cellulytic fungi and protozoa, various bacterial groups and methanogenic archaea, while bacteria characterize a typical gut microbiome of a monogastric.
The capabilities of a microbiome also relate to its composition. Each microorganism has different specific properties that complement and overlap with the characteristics of other microorganisms in a microbiome. This sum of sets of skills results, for example, in the performance, a microbiome can provide to the digestion of food and it determines the interactions between the microbiome and the host.
Such interactions can be desirable and undesirable effects. For example, on the one hand, the presence of microorganisms supports the development of the immune system and it establishes the acceptance of beneficial microorganisms. On the other hand, disorders in the composition of the microbiome or colonization of pathogenic microorganisms can lead to findings relevant for veterinary practice.
Concerning animal nutrition, microbiome-related issues are addressed through a combination of molecular biological, bioinformatical and statistical methods:
- Sequencing of marker genes from microorganisms; in cooperation with other institutes of the FLI
- Identification of microorganisms based on DNA sequences
- Analyzes of microbiome compositions based on classic ecological units
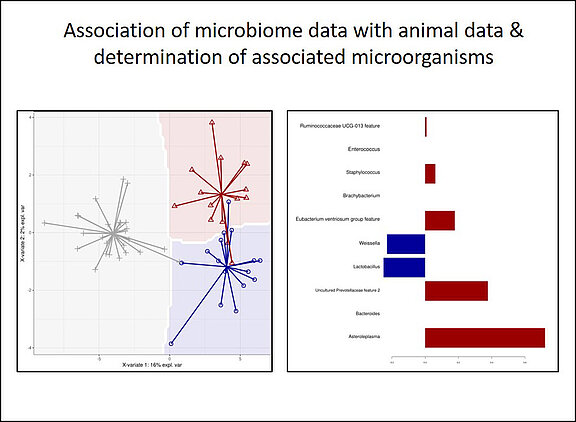
- Association of groups of microorganisms with observations from the host
Bioinformatics and modern computer science methods, which were developed in the field of machine learning are increasingly used in the observation and characterization of farm animals.
In comprehensive statistical procedures, for instance, results from methods for analyzing gene expressions are combined with data from various animal-specific analyzes. Such comprehensive analyzes of data provide an opportunity to get a relatively broad picture of the health status of an animal. In addition, findings from such comprehensive analyzes serve as the basis for training of machine learning models, which can be increasingly used for the automated description of animal conditions. It is a particular goal to prepare data from various sources, such as imaging processes, automatically in order to make it usable for automated machine learning methods.